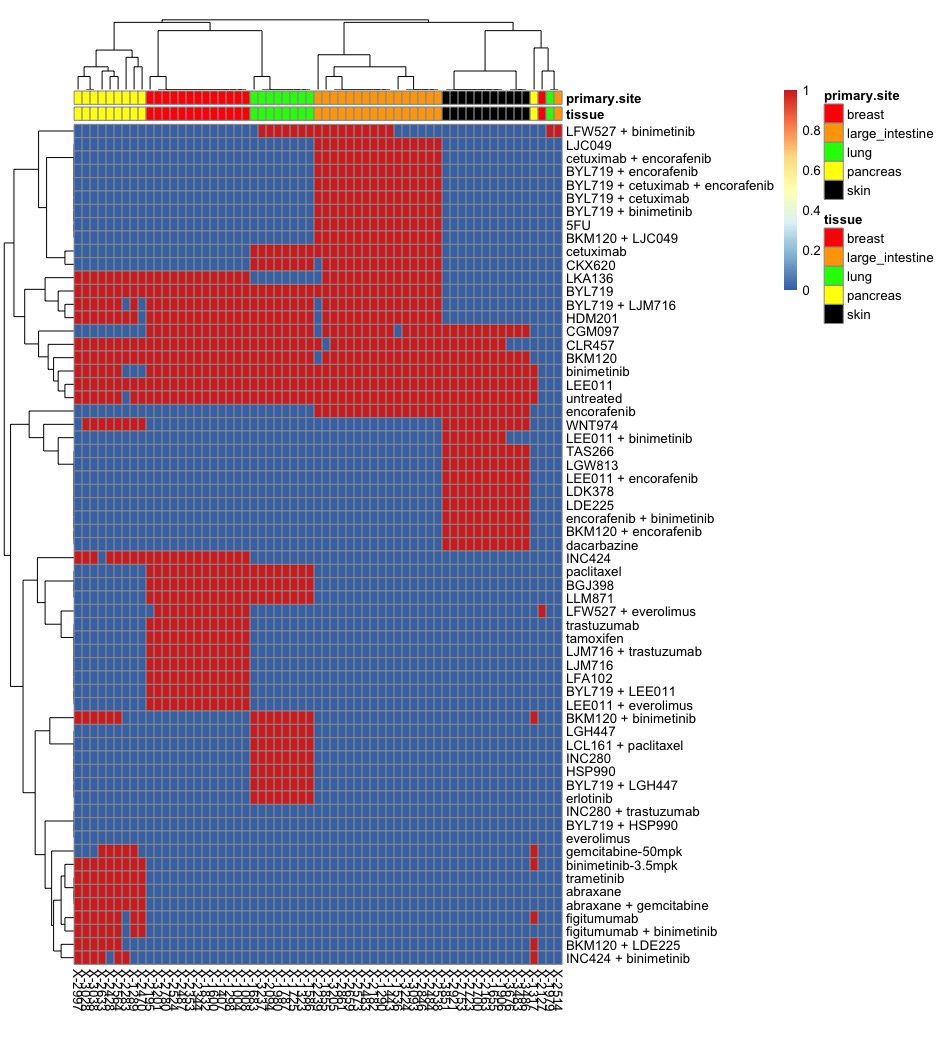
Overview
PDXGEM is a statistical bioinformatics and pharmacognomics pipeline to build a multi-gene expression model for predicting responses of cancer patients to anti-cancer therapeutics on the basis of data on mRNA expression profiles and drug-sensitivity of Patient-Derived tumor Xenograft models (PDXs). Please refer to the following preprint for more details.
- Kim Y. et al. (2020) PDXGEM: patient-derived tumor xenograft based gene expression model for predicting clinical response to anticancer therapy in cancer patients, BMC Bioinformatics, 21, 288 (2020). https://doi.org/10.1186/s12859-020-03633-z
Please click ‘PDXGEM(Guest)‘ menu to use PDXGEM web software without log-in. If you signed in, select ‘Run PDXGEM‘ menu.
PDXGEM process diagram
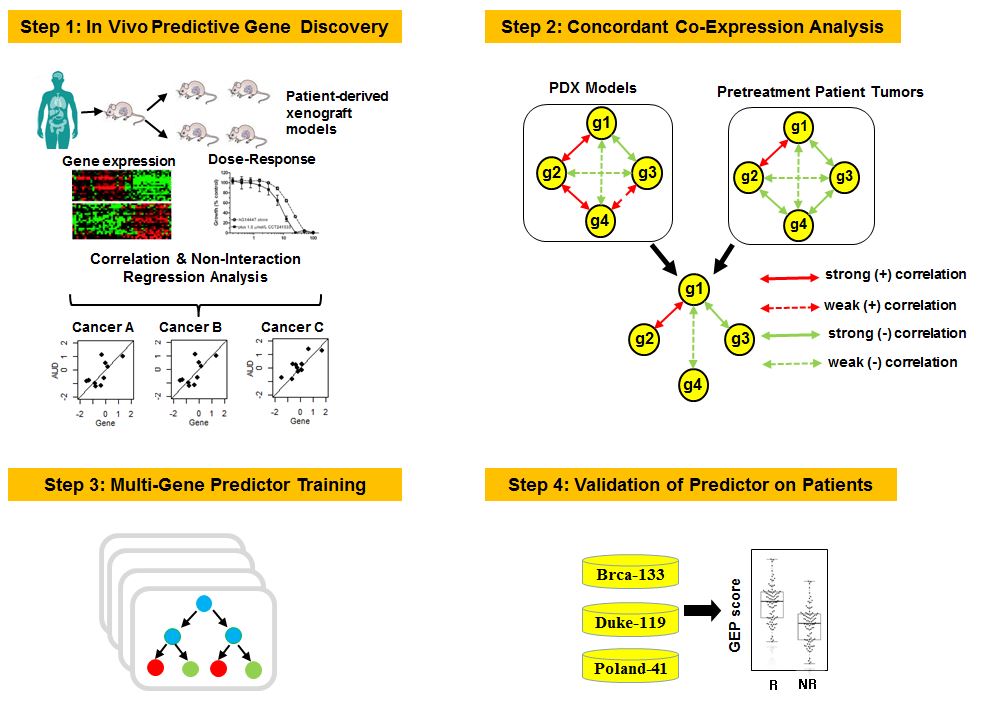
In brief, PDXGEM consists of four subsequent analysis steps, 1) in vivo drug senstivity biomarker discovery, 2) Concordant Co-Expression Analysis, 3) Multi-gene drug response prediction model training step, and 4) testing the performance of the prediction model.
Currently available PDXGEMs by cancer types and anti-cancer drugs