Tutorial
There are four input items to be selected at the menu panel of PDXGEM.
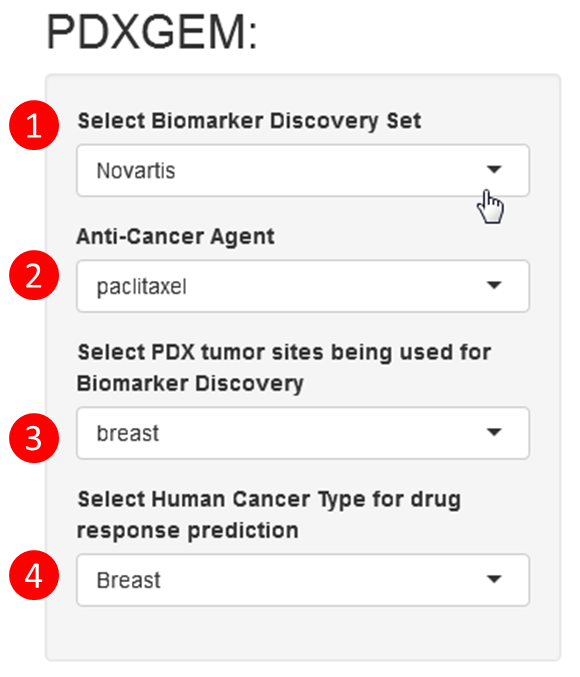
Step 1.1: Choose a Drug Sensitivity Biomarker Discovery set that is a PDX panel with available gene expression data and drug-sensitivity data. Novartis PDX panel data is currently available.
Step 1.2: Choose a anti-cancer drug of interest.
The distribution of drug sensitivity in terms of the percent of post-treatment tumor volume changes in PDX mouse models will be displayed in a vertical line plot and a boxplot as below.
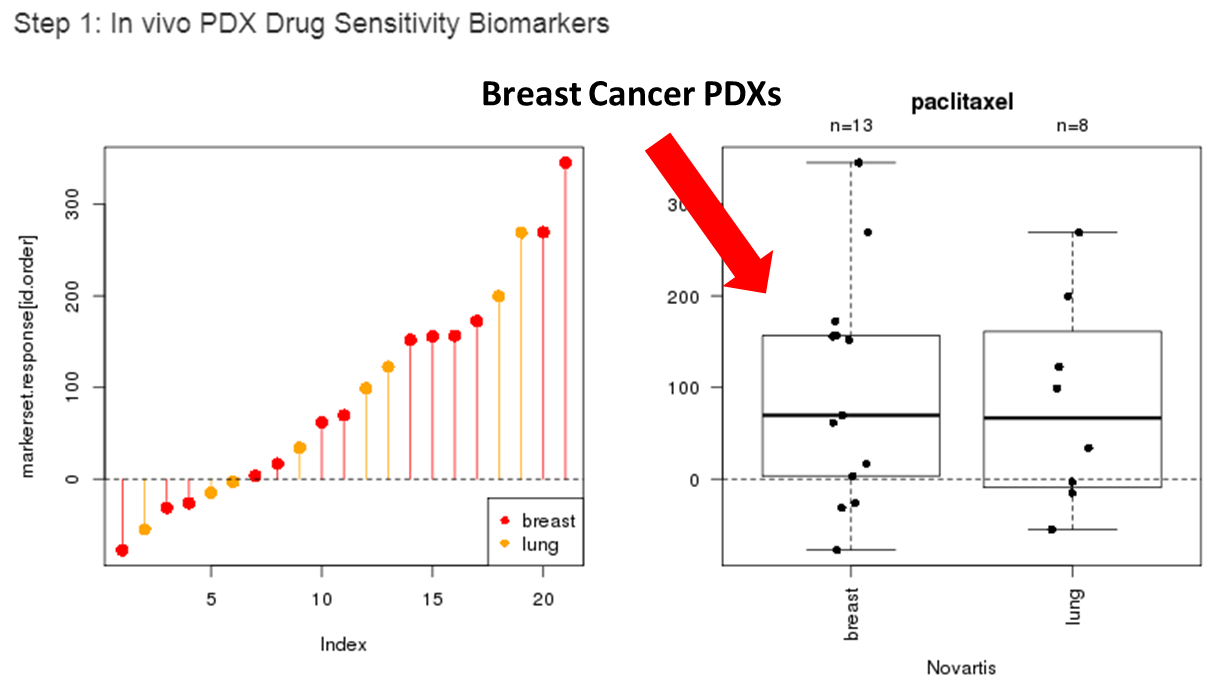
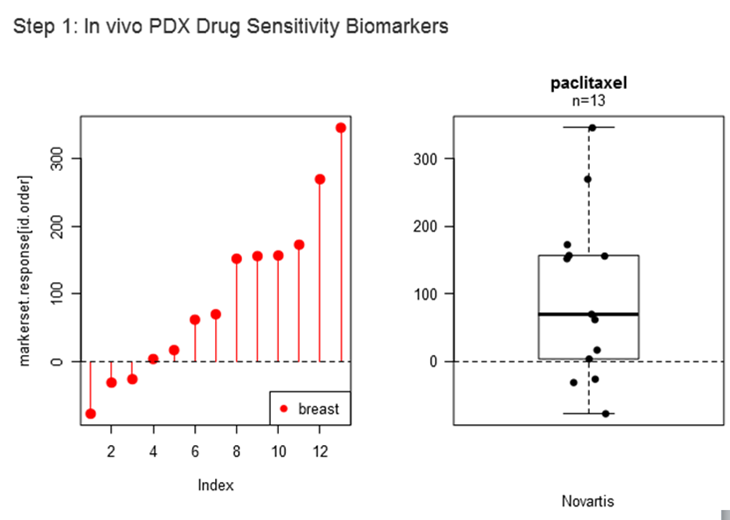
Step1.3: Choose all or a subset of PDX models that will be used to identify initial in vivo drug-sensitivity biomarkers.
The line and box-plots will be updated with the selected cancer type.
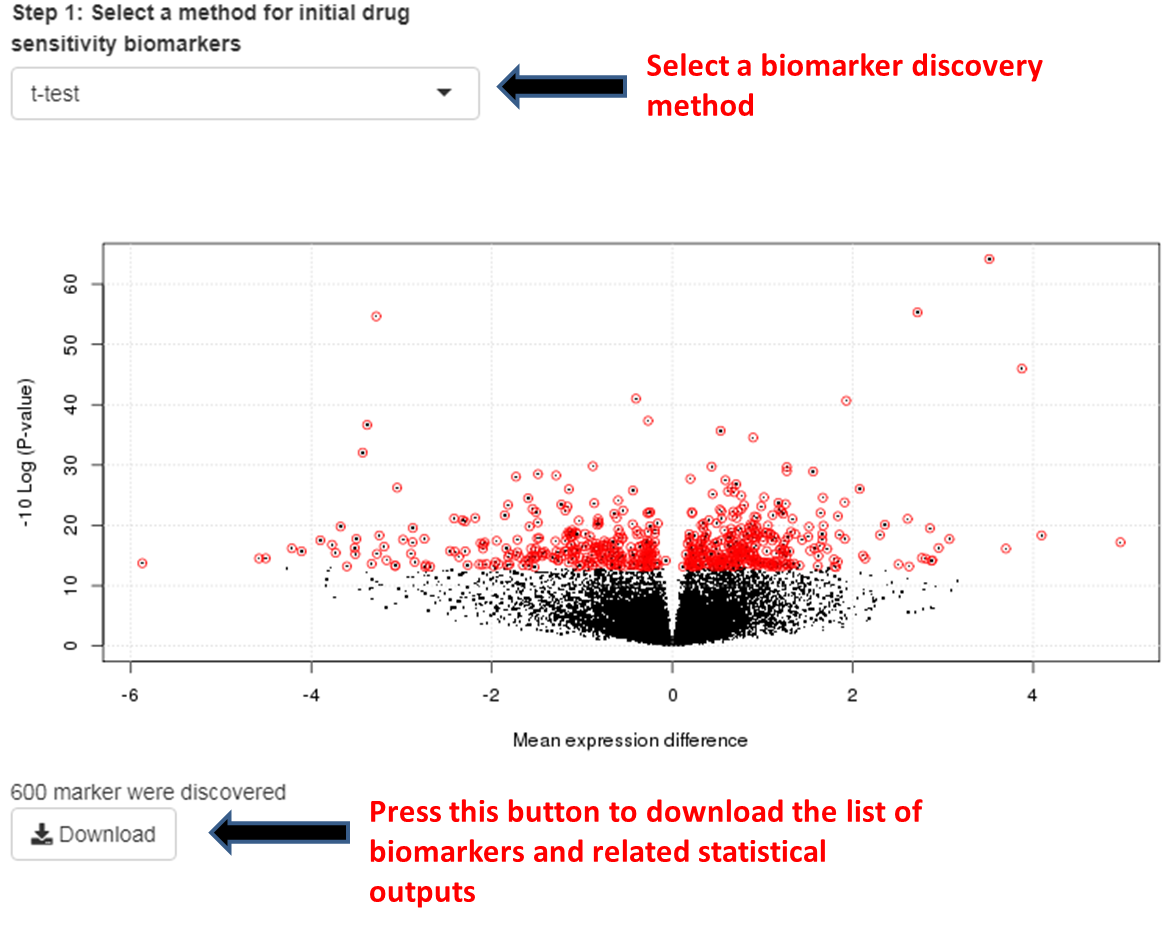
Step 1.4: Select a biomarker discovery method.
It will show a volcano plot of the degree of association between gene and drug-sensitivity. Red dots indicate the drug sensitivity biomarkers for the drug of interest. Press Download button below the volcano plot to download detailed results about the biomarkers.
Step 2.1 Choose a type of cancer of patients with which co-expression pattern of the drug sensitivity biomarkers in the PDX models will be compared in Comparative Co-Expression Analysis (CCEA).
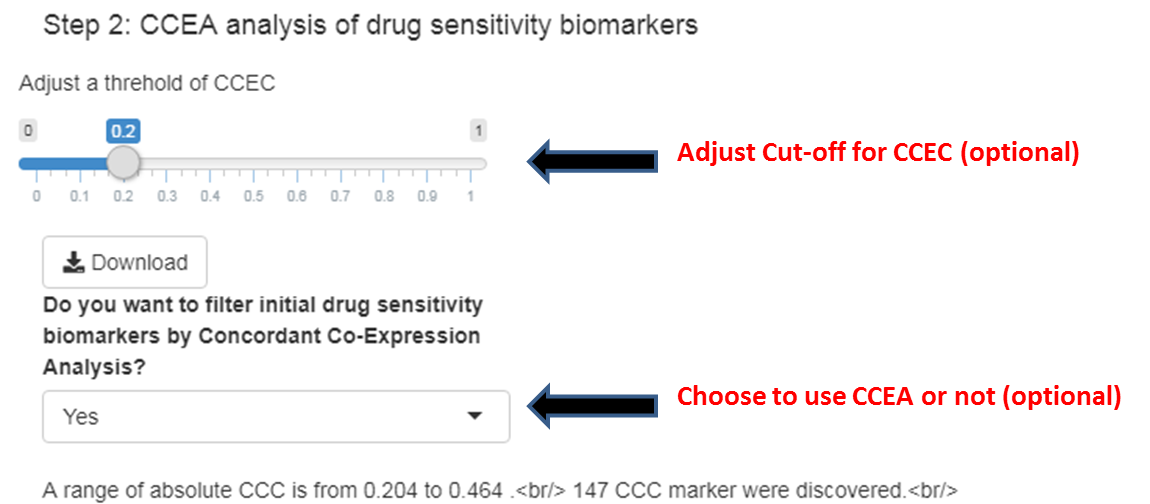
Step 2.2
You will see an output of CCEA between PDX models and representative cohort of patients with the selected cancer type. To evaluate the utility of CCEA, you can choose whether to use CCEA or adjust a cut-off for concordance-co-expression coefficient (CCEC).
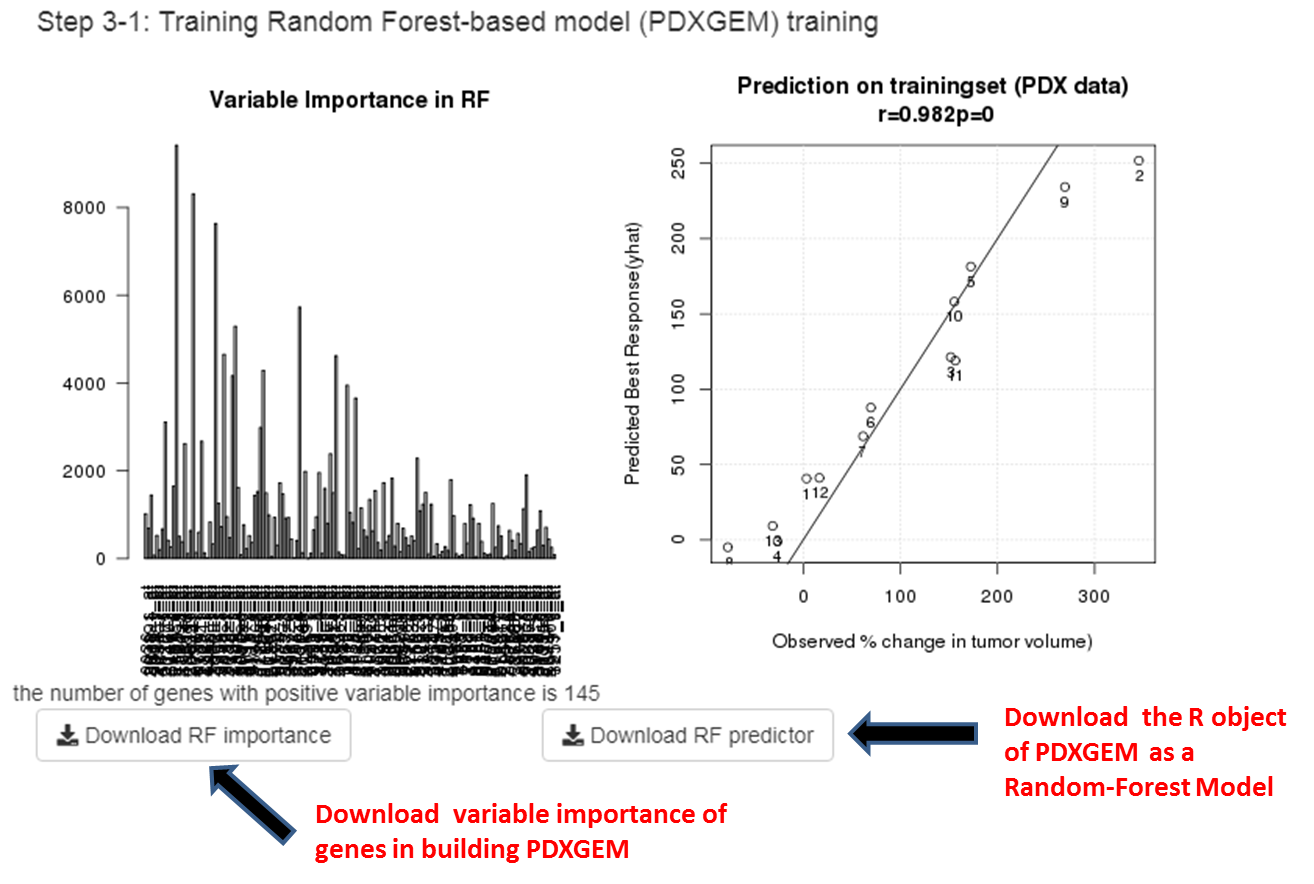
Step 3: Training PDXGEM to predict a drug response by Random Forest algorithm. A multi-gene based drug response prediction model (PDXGEM) will be build using Random Forest algorithm. You can download the list of genes used for training the PDXGEM as well as the R Random Forest object of the PDXGEM.
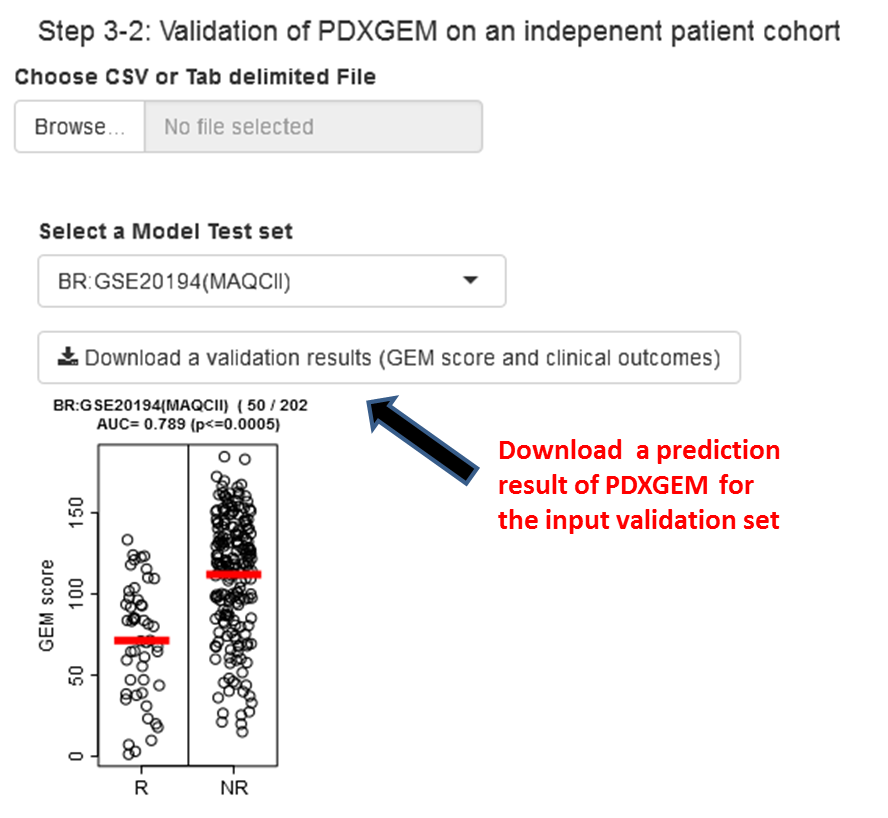
Step 4.1: Validate PDXGEM on the built-in test data set or user-uploaded dataset. Please see the input data format.
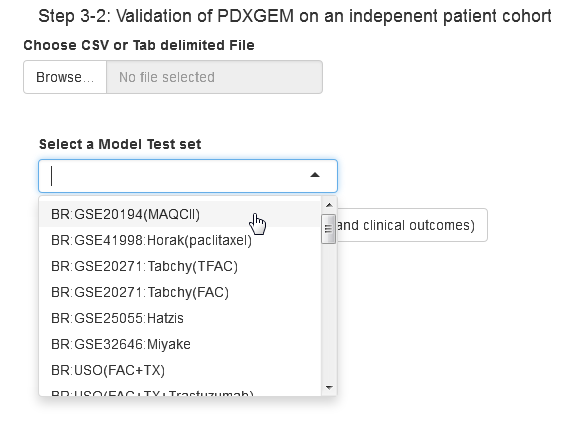
Step 4.2: Once validation data is selected, a strip-chart will be displayed as below. Users can download the final model and the prediction scores (tab-delimited text file).